Medicine is undergoing a fundamental transition towards leveraging molecular tools, driving a shift from reactive to proactive care. At the heart of this shift lies 5P medicine: preventive, predictive, precision, participatory, and population based approaches that acknowledge the intricate interplay between genetics, lifestyle, and environment.
Metabolomics, with its ability to track the behavior of small molecules, has a unique place in the 5P transformation, providing a measure of the impact of not only genetics, but also environment and lifestyle:
- Metabolomics enables preventive medicine through early, often reversible metabolic shifts that precede the onset of chronic disease
- Metabolomics enables predictive medicine through robust biomarker signatures already harnessed to foretell drug response
- Metabolomics enables precision medicine through stratification and metabotypes that group patients based on the molecular presentation of their disease,
- Metabolomics enables population based medicine by providing insight into environmental variance necessary to step into the post-genomic era
- Metabolomics enables participatory medicine through patient centric care including remote at-home sampling and actionable biochemical information.
This multifaceted perspective provided by metabolomics remains to be included in the routine toolkit of most medical practitioners, but when it does, it will drive a massive transformation in the way that we approach health.
Driving the adoption of metabolomics through 5P medicine
Here I introduce a simple but powerful model called Screen | Leverage | Translate. This 3-step framework explains how any scientist can harness the information contained in metabolomics and turn it into the solution to their specific 5P medicine problem.
If you are familiar with my book, the STORY principle, you’ll notice that Screen | Leverage | Translate follows a similar direction to the principle I describe there, planning an experiment, executing it, and excavating the gems out of your dataset before carving it into the answer you need for your research. Screen | Leverage | Translate is a streamlined version of the STORY principle, adapted to specific applications in 5P medicine.
Step 1 | Screen
This is the data acquisition step, where you’ll analyze the samples required for your experiment. Going back to the STORY principle, there is an unspoken understanding that by the time you reach sample measurement, you’ve already put in the work to determine your research question and how metabolomics will help answer it.
This doesn’t mean you know the results, rather that you’ve accounted for any factors that may predictably confound your findings. With a measure as sensitive to the environment as metabolomics, it is crucial to have these factors in check, in order to make the most of each study. Today, enough is known about these confounders to help you avoid interference by, e.g. sex, age, fasting time, diet, exposures, and more. If unsure, you can always refer back to the Safety Check step in my book or ask your metabolomics collaborators for support in planning your experiment.
A few questions to ask in preparation for this screening step relate to the method used for metabolomics measurement. Every method has benefits and limitations. Understanding the form of answer you need to your research question will turn certain benefits into requirements, and others into “nice-to-haves”. The method that provides your requirements without hindering the leveraging of the results is the one best suited for your study.
In our pathway spotlight below, I will describe an example focused on polyamine metabolites and their role in immunotherapy response.
Step 2 | Leverage
During your preparation step, you’ll also have planned the tools to be used to analyze and leverage your data. These days, with data and study design complexity increasing, the more automated the tool, the better.
Among omics, metabolomics is unique in the way that its datapoints connect to each other and to health. Unlike other omics, no direct line can be drawn between a given metabolite and a gene of origin. This makes the application of genomics-derived tools impossible, which is why the metabolomics community has had to develop its own data analysis tools.
Metabolites are most often grouped and analyzed around metabolic pathways. This work demands a deep knowledge of these mechanisms, that is condensed in databases like Wikipathways, Reactome and KEGG; databases that are constantly growing, as our knowledge of metabolism in different contexts expands.
Context is crucial.
Depending on the matrix you are studying (blood, urine, feces, tissue from an organ, cell model…), different metabolic pathways are at work – interacting, influencing each other, balancing each other.
The optimal way to leverage metabolomics is intricately linked to the context of the study. Species, matrix, disease context, study design – when combined, these factors make each experiment unique. But knowledge from previous experiments is a great way to start leveraging your own. I’ll introduce the one we have developed in our software in our pathway spotlight example.
Step 3 | Translate
Translation is where every omic meets its end application; where the well-oiled machine of data acquisition needs to slow down and level with what awaits it on the other side.
For applications where a routine solution is the expected outcome, translation is just the creative part before another well-oiled machine is created; e.g. a biomarker panel for routine disease screening, a biomarker for drug response prediction, a bioinformatics pipeline combining metabolomics with other omics and medical endpoints.
Seen through this lens, translating metabolomics is the thrilling part. It is the moment of truth, where the results of the previous steps are put to the test: will they hold up in a follow-up study? Because indeed, after crafting the solution to your original research question, you’ll need to verify its power in one or more validation studies.
This is when research begins to bring value to society; how we truly leverage knowledge to improve lives. Three years after its publication, I still refer to the paper by Tintelnot et al. on pancreatic cancer patient response to chemotherapy. Using metabolomics, this team identified a microbial metabolite, 3-indole acetic acid (3-IAA), as higher at baseline in patients who responded to treatment. Many studies end there, but not this time. What followed was a set of experiments in an animal model of pancreatic cancer, testing the impact of the patients’ microbiomes in fecal matter transplantation (FMT) experiments, an evaluation of the effect of direct supplementation with 3-IAA and other metabolites, and painting the picture of how 3-IAA modulated the immune system of the mice to improve chemotherapy response.
These are the kinds of studies I love to read: where omics are leveraged to start crafting the solutions to a specific problem. This is why translation is called “translation”. It takes work and creativity to transform the tabular results provided by omics into the life-sized solutions needed in the clinics.
In the pathway spotlight below, I explore how metabolites, specifically polyamines, may bring similar solutions for immunotherapy response. My goal here is to walk you through bits of published science to understand how you could leverage and translate your next metabolomics experiment to accelerate its adoption in the future medical landscape.
Pathway spotlight | Polyamine acetylation as a driver of immunotherapy response
Metabolomics profiling creates a broad picture of the metabolome. Depending on the level of expertise of scientists, they may enter this step with a fully blank mind or with an idea of what they think they will see. This second case is ideal, as it can also use some of the results as a form of positive control for the phenotypes that they are studying.
Here we will work under the assumption that broad metabolic profiling identified differences in polyamines in a cohort of patients receiving immunotherapy treatment for cancer. Since this pathway is relevant in several types of cancer, I will not specify here and rather detail the disease context each time that a study is discussed in this spotlight
Step 1 | Screen
In a study of patients undergoing anti CD19 CAR T-cell therapy for relapsed or refractory large B cell lymphoma, Fahrmann et al. found that acetylated polyamines, specifically acetylspermidine and diacetylspermidine, were elevated in non‑responders and strongly linked to poor treatment durability (Fahrmann et al. 2022).
The first thing any scientist would do is to research what these metabolites are known for.
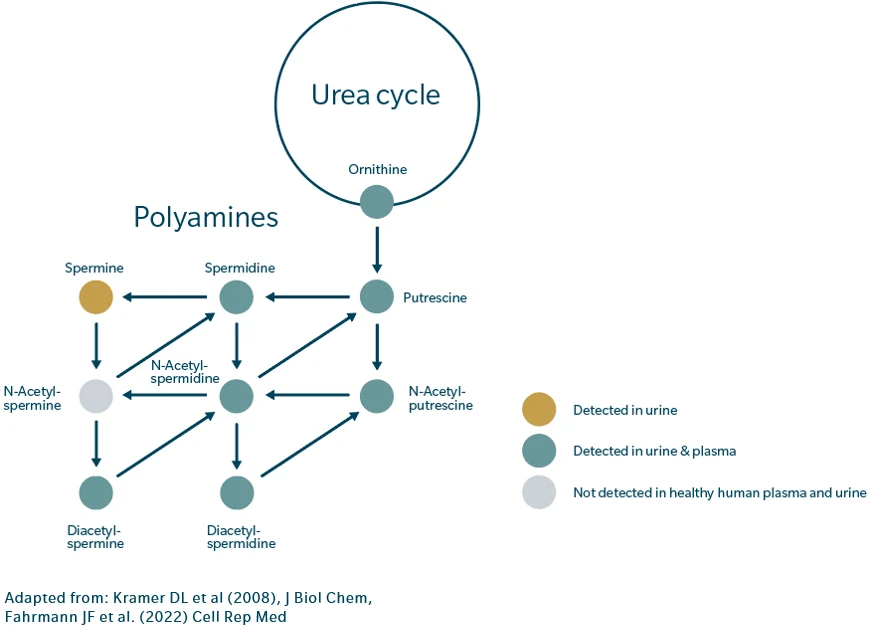
Polyamines – putrescine, spermidine, and spermine – are essential molecules for cell growth, gene regulation, and immune function. Cancer cells exploit this pathway aggressively. A key regulator of this system is spermidine/spermine N¹ acetyltransferase (SSAT), the enzyme responsible for polyamine acetylation and export.
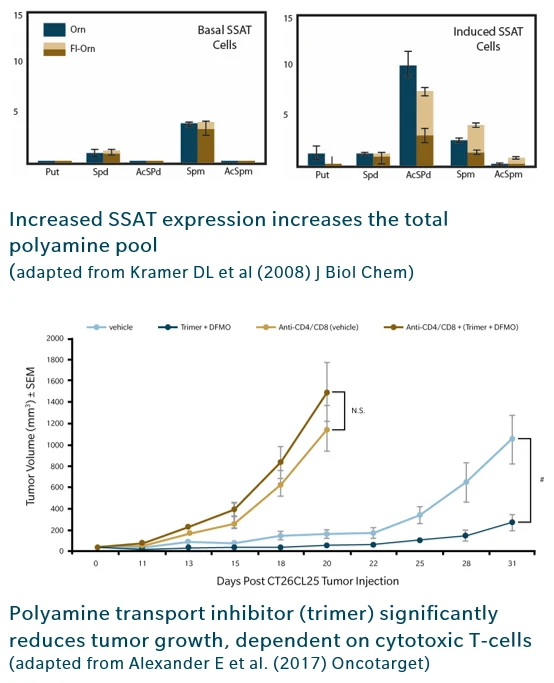
Mechanistic investigations show that overexpression of SSAT dramatically increases flux through the polyamine pathway, triggering both acetylation and compensatory upregulation of synthesis. This “futile cycle” results in a large, constantly replenished polyamine pool (Kramer et al. 2008). In addition, acetylated polyamines accumulate in several cancers, including breast, prostate, and lung tumors, contributing to an immunosuppressive microenvironment that blunts therapeutic efficacy. (Thomas & Thomas, 2007, Kramer et al. 2008)
This information is valuable and suggests that these metabolic differences may link to differences in treatment response. However, literature research can be a long and confusing endeavor. This is why the whole community strives to automate leveraging and interpreting metabolomics results. In the following step, we’ll review how MetaboINDICATOR can facilitate the first steps of this work.
Step 2 | Leverage
Enough data has been generated by now to inform on the expected metabolic profile of at least part of an experiment and better understand its context. This is what MetaboINDICATOR was designed for. This module of the WebIDQ software contains hundreds of sums and ratios of metabolites hand-picked from the literature and distributed across disease- and mechanism-related categories.
The most open-ended way to use this module is to compute any indicator for which the metabolites are quantified in the sample. This is my recommendation to maximize the chance to learn from other fields; although it is also possible to select only the groups of indicators of interest to the study, for example indicators related to cancer.
After statistical analysis of these new datapoints, data interpretation can begin. MetaboINDICATOR provides references to the literature where each sum or ratio has been previously described, thus immediately expanding the perspective for application.
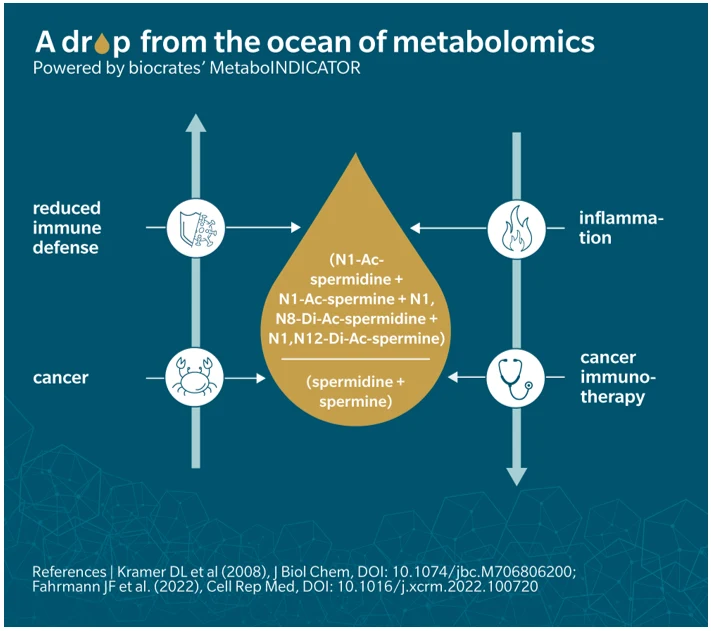
Let’s take the example of the ratio of acetylated to non-acetylated polyamines. Experimental work in cancer models shows that this ratio correlates strongly with SSAT enzyme expression, providing a functional handle on acetylation dynamics (Kramer et al. 2008). While the ratio of product to substrate can be used as a proxi for enzymatic activity, interpretation based on the biological context is key. What is most interesting here though is the correlation of these metabolite levels with immunotherapy response.
For someone with little to no experience in metabolomics, having this automated access to a relevant indicator and literature attached to it can save hours of literature research and put them directly on the right track for their interpretation, where the precise context of their research is going to be a strong driver of their thinking process towards translation.
Step 3 | Translate
For this step, I’ll go back to the 5P medicine concept and pick examples for predictive and precision medicine applications of metabolomics.
Predictive Medicine: Forecasting immunotherapy success
Recent clinical research demonstrates that polyamine related metabolites are predictive of patient response. Elevated plasma acetylated polyamines are associated with poor response to CAR T-cell therapy in relapsed/refractory large B cell lymphoma. A 6 marker metabolite panel (including acetylspermidine and diacetylspermidine) was validated across cohorts as a predictor of non durable response (Fahrmann et al. 2022). This was the outcome of the very first study I introduced in step 1.
Identifying a biomarker signature is one of the best-known applications of metabolomics, and with strong results in validation cohorts, this can lead to deep impact in how medicine will be practiced in the field. But I also wanted to include an example focused on new drug therapy development.
Precision Medicine: Polyamine blockade therapy
Targeting polyamine metabolism doesn’t stop at prediction; it also offers therapeutic opportunity.
A landmark study introduced polyamine blockade therapy, a dual strategy combining inhibition of polyamine synthesis (DFMO) with a polyamine transport inhibitor (Trimer PTI). Polyamine blockade therapy significantly reduced tumor growth more effectively than either inhibitor alone (Alexander et al. 2017). The anti tumor effect is T cell dependent, featuring increased cytotoxic CD8⁺ T cells and reduced immunosuppressive cell populations. By reversing the immunosuppressive effects of high polyamine levels, this approach exemplifies how metabolomics informed targets can directly shape therapeutic innovation.
This pathway spotlight illustrates how metabolomics, guided by the Screen | Leverage | Translate framework, enables deep biological insight and clinical translation. Polyamine metabolism is just one pathway among many, yet its story captures the essence of what 5P medicine strives for: using precise biochemical information to create predictive, preventive, and precise health solutions, from individual to population health.
As you explore your own datasets, I encourage you to adopt this structured approach. In particular, don’t stop at the screening step. Dig further. Leverage your data to gain a broader understanding of the metabolic changes in your system, and devise the next experiments that will yield the truly transformative solutions to transform medicine.




